 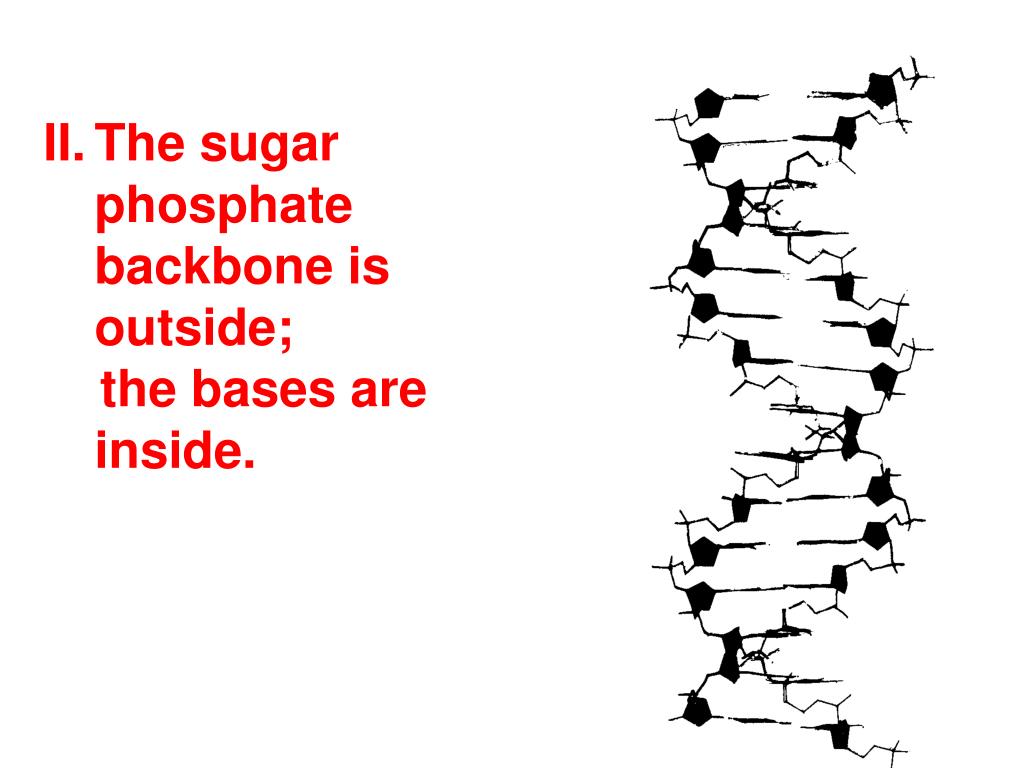 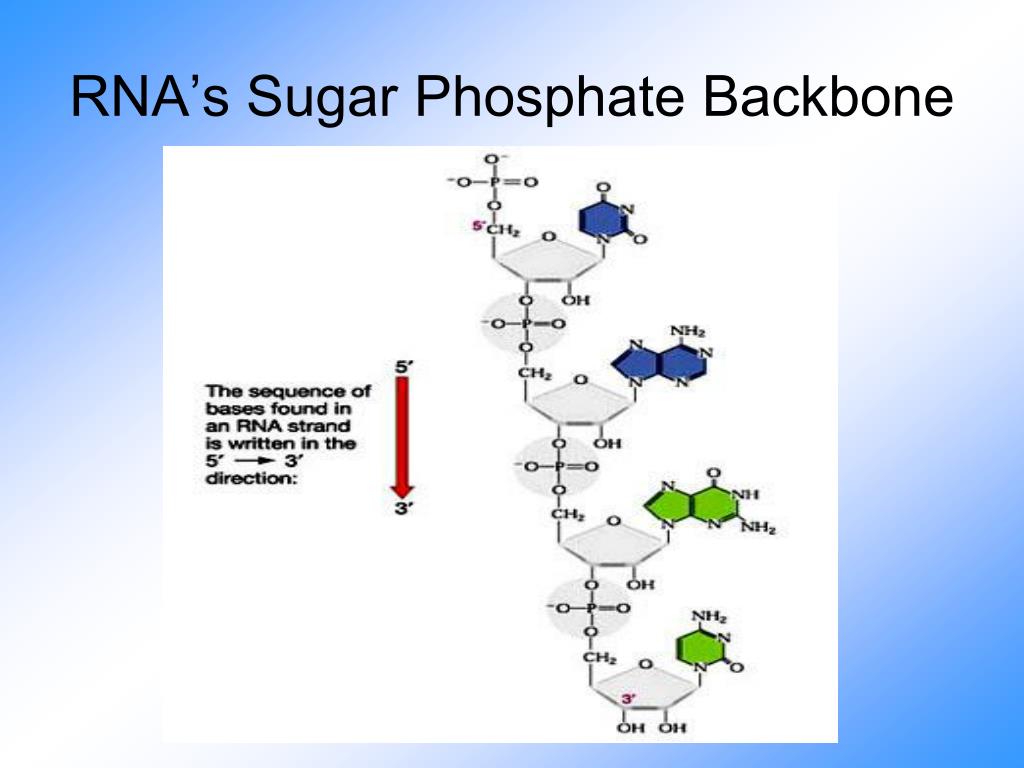
So in a polymer of multiple nucleotides (DNA, RNA), the repeating monomer unit contains two phosphoester bonds, on "top" (5') and "below" (3') the sugar. Additionally, two nucleotides are connected by one phosphoester bond (between a phosphate O and sugar 3'-C). Įxisting ribozymes can also be altered in a way that changes the reaction(s) that the ribozyme catalyzes.Each nucleotide contains one phosphoester bond (between a phosphate O and sugar 5'-C). Splicing ribozymes catalyze RNA splicing, removing a section of RNA that contains a mutation and replacing it with well-functioning RNA. In addition to inhibiting gene expression, splicing ribozymes can be used to repair damaged or defective RNA. These ribozymes can be designed to prevent expression of a particular gene, for example. Hammerhead ribozymes, in particular, can be designed such that they will cleave a desired RNA. Applications include the use of ribozymes in gene therapy to control gene expression in bacteria and eukaryotes, and to inhibit viral replication. Researchers are developing and using various applications for RNA hydrolysis that can be carried out in a controlled way. Large ribozymes, such as Group I introns, Group II introns, and RNase P, catalyze splicing and other post-trascriptional modifications during mRNA processing, using the cleavage mechanism described above. Examples of such ribozymes include the hammerhead ribozyme, the Hepatitis Delta Virus (HDV) ribozyme, and the hairpin ribozyme. Ī category of ribozymes called small ribonucleolytic ribozymes enhances the spontaneity of the cleavage of their own RNA using acid-base catalysis. A lysine residue, also in the active site of RNase A, stabilizes the negatively charged oxygen atoms in the transition state. Certain histidine residues in the active site act as bases to remove protons from 2’ hydroxyls of ribose sugars, while others act as acids to donate protons to the 5’ oxygen of adjacent riboses to make them better leaving groups. RNase A contains histidine in its active site, and uses it to accomplish acid-base catalysis and cleavage of RNA. One such enzyme is Ribonuclease A (RNase A), a protein enzyme. Several different enzymes catalyze cleavage at specific sites on an RNA molecule. In these instances, cleavage is done using catalytic enzymes. When the RNA is double-stranded or involved in nucleotide base pairing, it is more stable and spontaneous cleavage is significantly less likely. This deprotonation makes the reaction base-catalyzed and increases spontaneity of the reaction. Auto-hydrolysis or self-cleavage reactions take place in basic solutions, where free hydroxide ions in solution can easily deprotonate the 2’ OH of the ribose. Spontaneous cleavage in an RNA molecule is much more likely to occur when it is single-stranded. This process is known as an auto-hydrolysis or a self-cleavage reaction. The hydrolysis or cleavage of RNA can occur spontaneously, without the presence of a catalyst or enzyme. (This mechanism is also referred to as RNA cleavage.) This produces a 2’,3’-cyclic phosphate that can then yield either a 2’- or a 3’-nucleotide when hydrolyzed. The phosphorus then detaches from the oxygen connecting it to the adjacent sugar, resulting in ester cleavage of the RNA backbone. There is a transition state (shown above), where the phosphorus is bonded to five oxygen atoms. RNA hydrolysis occurs when the deprotonated 2’ OH of the ribose, acting as a nucleophile, attacks the adjacent phosphorus in the phosphodiester bond of the sugar-phosphate backbone of the RNA. 4) The 2′,3′-cyclic phosphate group (in step 3) hydrolyzes to either the 2′ or 3′ phosphate. 3) Phosphodiester bond is broken, cleaving the RNA backbone. 1) Base-catalyzed deprotonation of the 2′-OH group, enabling the deprotonated 2′ hydroxyl's nucleophilic attack on the adjacent phosphorus. Mechanism of base catalyzed RNA hydrolysis. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
